Sitemap
A list of all the posts and pages found on the site. For you robots out there is an XML version available for digesting as well.
Pages
Posts
Future Blog Post
Published:
This post will show up by default. To disable scheduling of future posts, edit config.yml and set future: false.
Blog Post number 4
Published:
This is a sample blog post. Lorem ipsum I can’t remember the rest of lorem ipsum and don’t have an internet connection right now. Testing testing testing this blog post. Blog posts are cool.
Blog Post number 3
Published:
This is a sample blog post. Lorem ipsum I can’t remember the rest of lorem ipsum and don’t have an internet connection right now. Testing testing testing this blog post. Blog posts are cool.
Blog Post number 2
Published:
This is a sample blog post. Lorem ipsum I can’t remember the rest of lorem ipsum and don’t have an internet connection right now. Testing testing testing this blog post. Blog posts are cool.
Blog Post number 1
Published:
This is a sample blog post. Lorem ipsum I can’t remember the rest of lorem ipsum and don’t have an internet connection right now. Testing testing testing this blog post. Blog posts are cool.
portfolio
Portfolio item number 1
Short description of portfolio item number 1
Portfolio item number 2
Short description of portfolio item number 2 
publications
Recursively constructing analytic expressions for equilibrium distributions of stochastic biochemical reaction networks
Published in J. Royal Soc. Interface, 2017
Text… 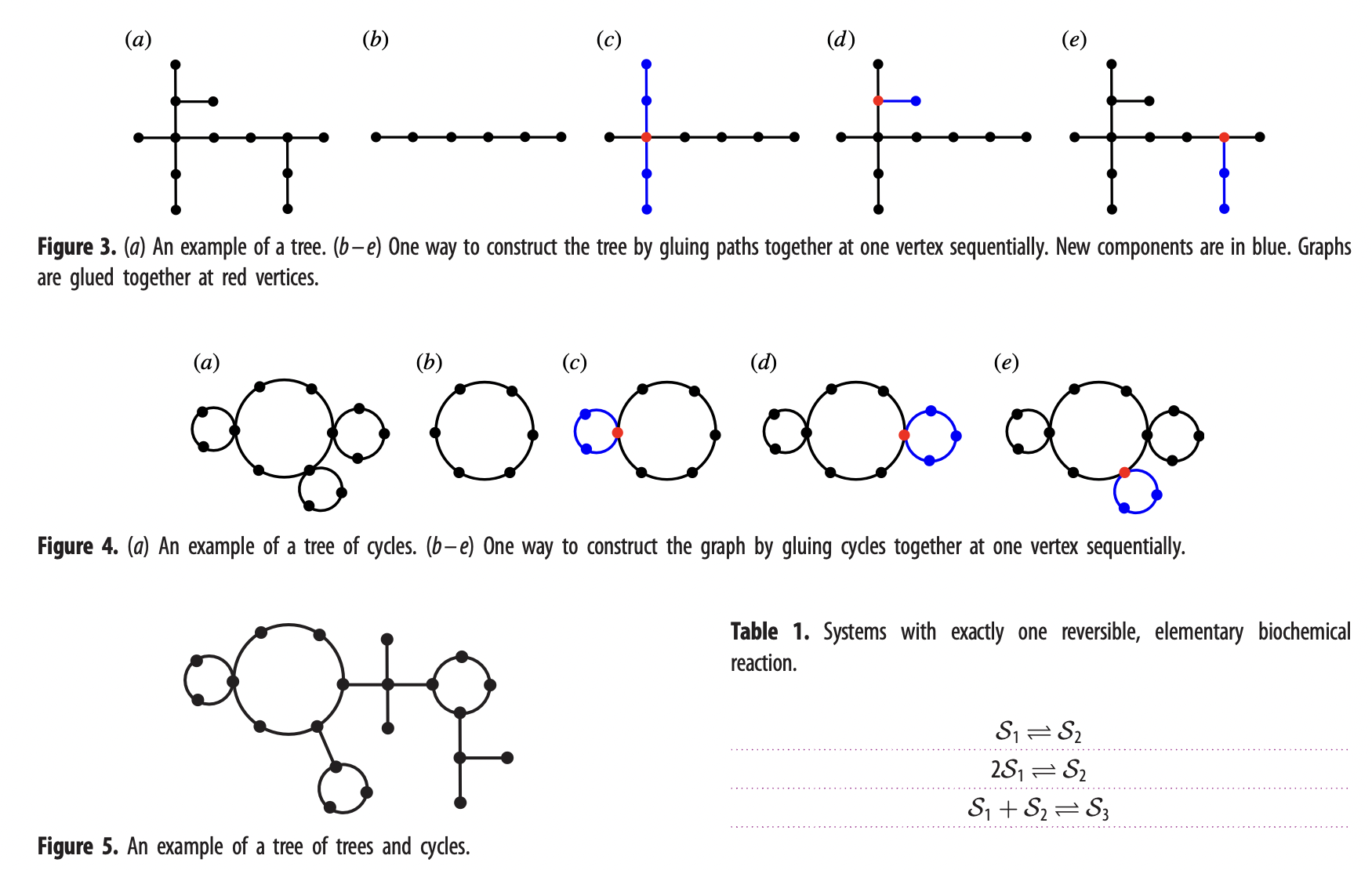
Recommended citation: Meng X. Flora, Baetica Ania-Ariadna, Singhal Vipul and Murray Richard M. (2017) "Recursively constructing analytic expressions for equilibrium distributions of stochastic biochemical reaction networks" J. R. Soc. Interface.(14) 20170157. http://doi.org/10.1098/rsif.2017.0157
Transforming Data Across Environments Despite Structural Non-Identifiability
Published in American Control Conference, 2019
We present a framework for batch correcting genetic circuit data across environments, and show that parameter structural non-identifiability need not hinder this goal. We give parameter consistency conditions under which we can perform such correction despite the parameters not being identifiable.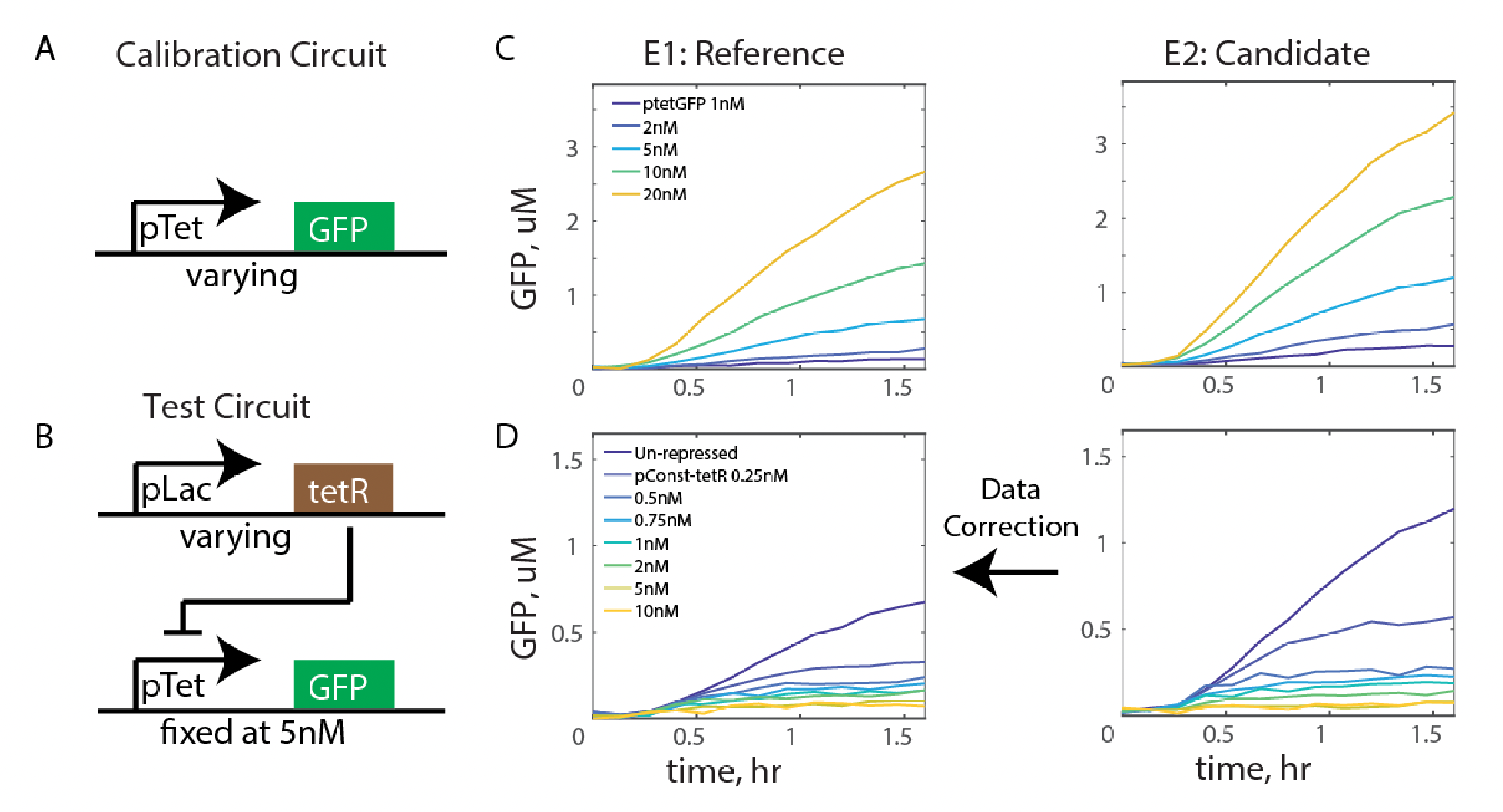
Recommended citation: V. Singhal and R. M. Murray (2019). "Transforming Data Across Environments Despite Structural Non-Identifiability." American Control Conference (ACC), Philadelphia, PA, USA, 2019 ,pp. 5639-5646, doi: 10.23919/ACC.2019.8814953.
A MATLAB toolbox for modeling genetic circuits in cell-free systems
Published in OUP Synthetic Biology, 2021
Txtlsim is a toolbox for simulating cell free reactions using mass action kinetics I this paper, we show how models of subsystems of a circuit can be individually characterized, and composed into the full system, whose behavior can be accurately predicted.
Recommended citation: Singhal et al. "A MATLAB toolbox for modeling genetic circuits in cell-free systems." Synthetic Biology. Volume 6, Issue 1, 2021, ysab007. https://doi.org/10.1093/synbio/ysab007
DUBStepR is a scalable correlation-based feature selection method for accurately clustering single-cell data
Published in Nature Communications, 2021
DUBStepR is a feature selection method that relies on the intuition of finding the most informative features given the set of features already identified.
Recommended citation: Ranjan, B. et al. "DUBStepR is a scalable correlation-based feature selection method for accurately clustering single-cell data." Nat. Commun.. 12, 5849 (2021). https://doi.org/10.1038/s41467-021-26085-2
BANKSY unifies cell typing and tissue domain segmentation for scalable spatial omics data analysis
Published in Nature Genetics, 2024
BANKSY is an algorithm with R and Python implementations that identifies both cell types and tissue domains from spatially-resolved -omics data by incorporating spatial kernels capturing microenvironmental information, applicable to a range of spatially-resolved technologies, and scalable to millions of cells.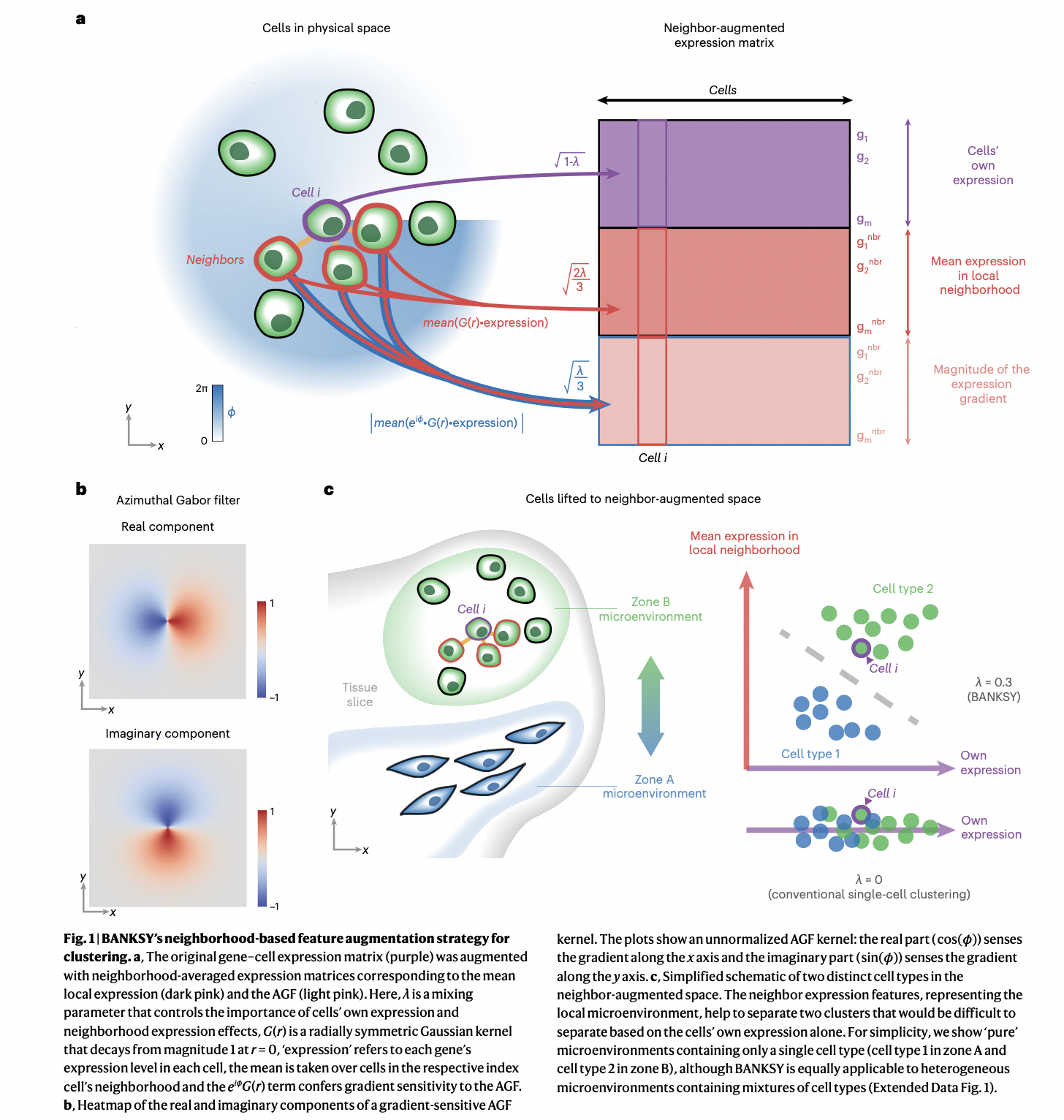
Recommended citation: Singhal, V., Chou, N., Lee, J. et al. (2024). "BANKSY unifies cell typing and tissue domain segmentation for scalable spatial omics data analysis." Nat Genet. https://www.nature.com/articles/s41588-024-01664-3
talks
Transforming Data Across Environments Despite Structural Non-Identifiability
Published:
In this work, we developed a theoretical framework for batch correction in cell-free systems in the presence of parameter (structural) non-identifiability. After formally defining the problem, we develop a set of parameter consistency conditions which guarantee the accuracy of the batch correction methodology despite parameter uncertainty. We also demonstrated it on real genetic circuit behavior in two batches of cell free extracts.
BANKSY unifies cell typing and tissue domain segmentation for scalable spatial omics data analysis
Published:
teaching
Design Principles for Genetic Circuits (2014)
Course, Broad 200, 2014
I TA’d a course called Design Principles for Genetic Circuits in 2014 and 2015. In 2014, the course was taught by Richard Murray and Michael Elowitz. You can access the link to the course webpage for 2014 here
Design Principles for Genetic Circuits (2015)
Course, Broad 100, 2015
In 2015, I TA’d Caltech’s BE 150 course. We covered design principles for genetic circuits, and various simulation and modeling frameworks. You can access the web archive link to the course webpage here
Mentorship (undergraduate and masters students)
Mentorship, Caltech and GIS, 2020
Over the years, I have had the opportunity to mentor and work with various students. Here is a list of their projects.
Spectral Graph Convolution Nerual Networks
Seminar Course, Bioinformatics Institute (BII), 2022
Between August 2022 and Feb 2023, Dr Hwee Kuan Lee at the Bioinformatics Institute taught a course on graph neural networks to researchers at A*STAR. I helped teach a submodule within this course on the graph fourier transform and spectral graph convolution neural networks. You can find some of the slides here.
Riemannian Geometry
Recitation / Tutorials, Bioinformatics Institute (BII), 2023
Between February and November 2023, I assisted Dr Hwee Kuan Lee in teaching a course on differential geometry to AI researchers in Singapore’s Agency for Science Technology and Research.
