BANKSY unifies cell typing and tissue domain segmentation for scalable spatial omics data analysis
Published in Nature Genetics, 2024
Spatial omics data involve measurements of both the molecular profiles (such as RNA or protein counts) and the spatial locations of cells. This allows us to measure the arrangement of individual cell types in stereotypical spatial patterns, which is a fundamental property of solid tissues.
The data resulting from these technologoes can be clustered to define both cell types and tissue domains. We have developed an algorithm called BANKSY, which unifies these two spatial clustering problems under a single biologically motivated framework. Cells are embedded in a product space of their own and the local neighborhood transcriptomes, representing cell state and microenvironment respectively.
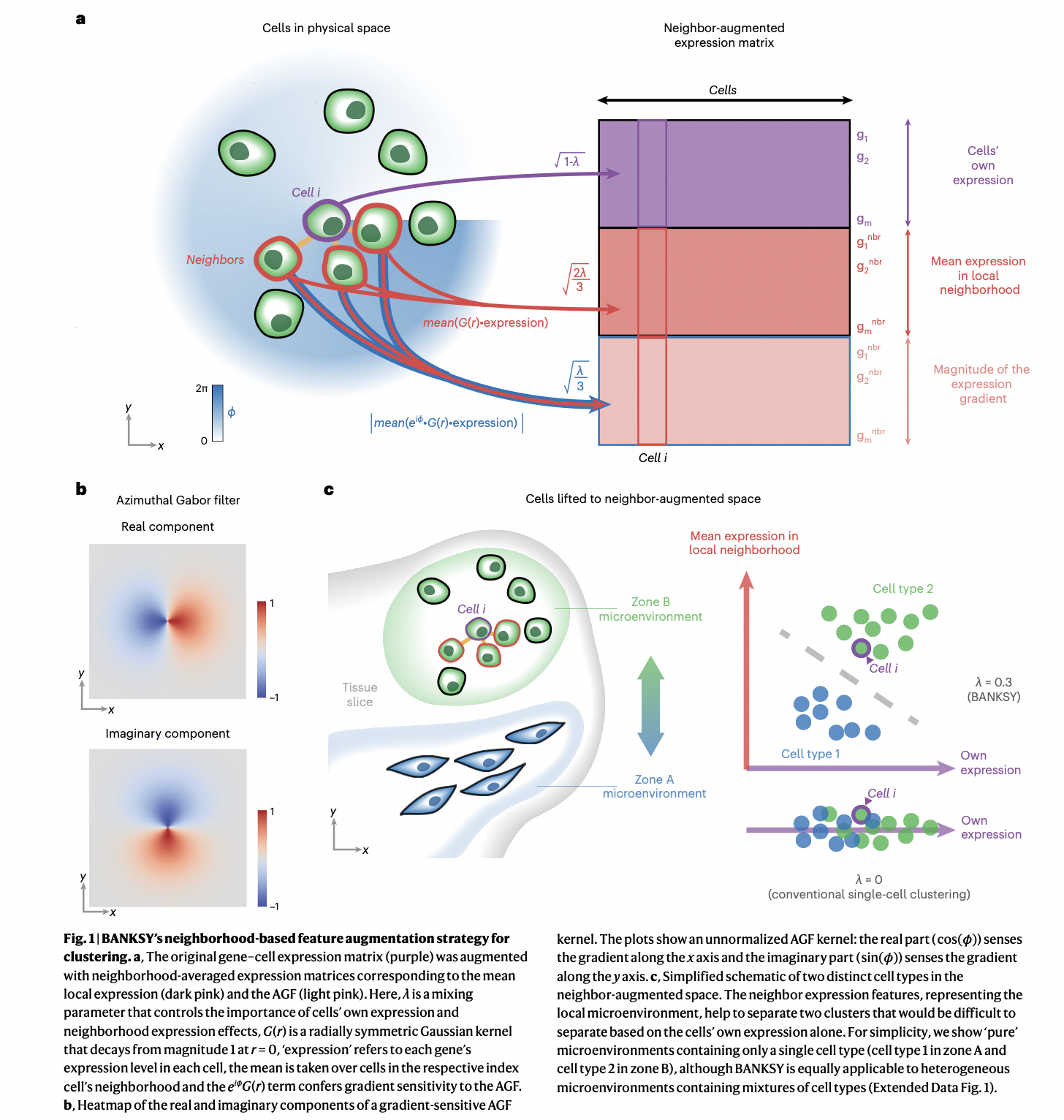
The neighbourhood is encoded using the local mean transcriptome, and local neighbourhood gradients computed using the azimuthal Gabor filter (AGF), weighted by a mixing parameter, lambda. The AGF is like the traditional plane wave Gabor filter used to quantify texture in image processing, but computed in the azumuthal (polar) coordinates around each cell, and modulated with a Gaussian waveform. You can read a summary article contextualizing things here.
Recommended citation: Singhal, V., Chou, N., Lee, J. et al. (2024). "BANKSY unifies cell typing and tissue domain segmentation for scalable spatial omics data analysis." Nat Genet. https://www.nature.com/articles/s41588-024-01664-3
